Viewing and Downloading Results
Subtopics
Query Results
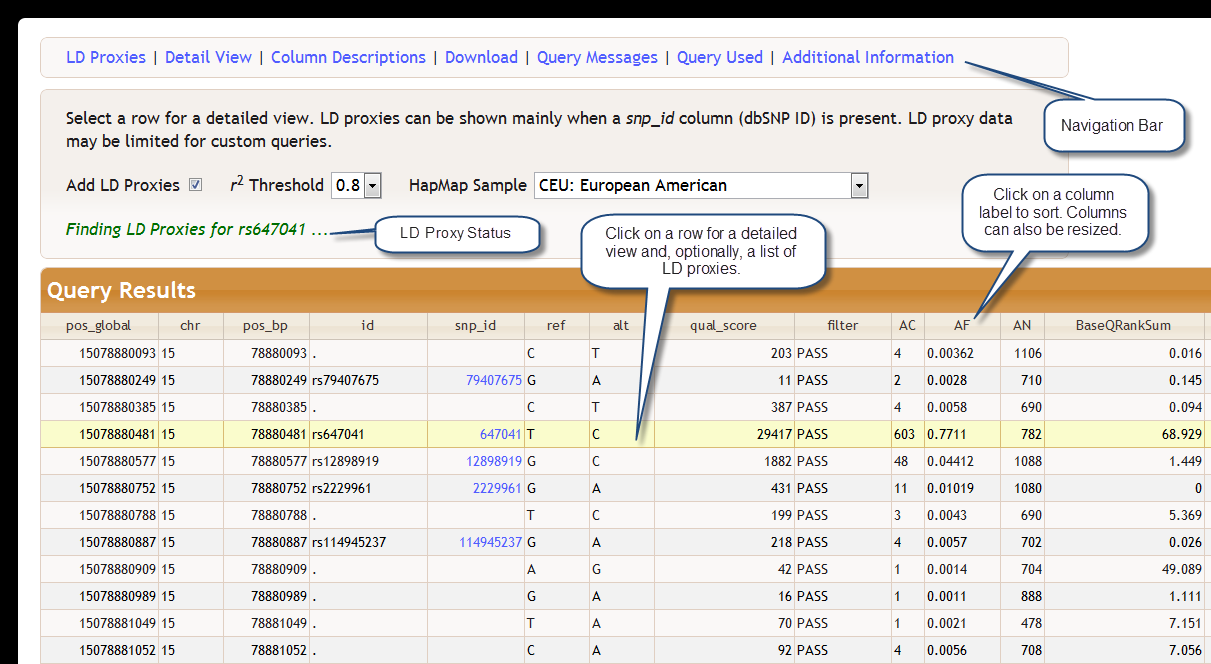
After a query is executed a new section appears at the bottom of your browser. Query results are viewed in an interactive table with a direct connection to our database server for powerful sorting and navigation features that can be viably used even when there are over one million rows.
Table Navigation and Column Descriptions
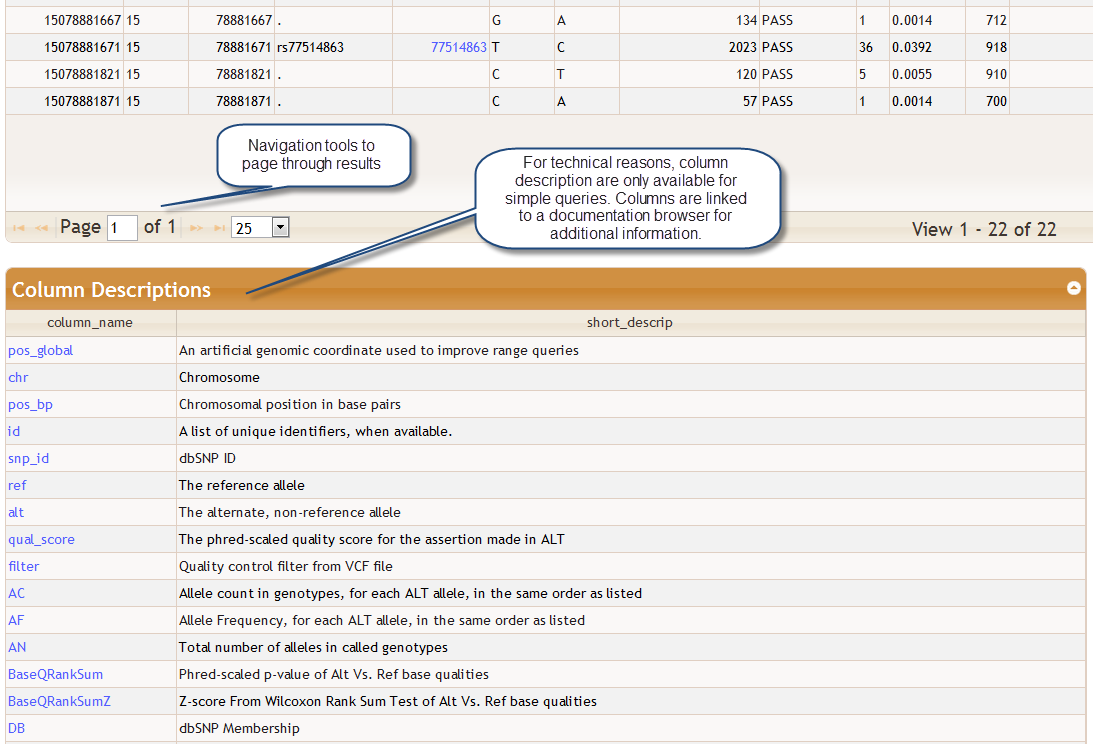
There is a toolbar at the bottom of the query results table that allows you navigate the table and change the number of rows shown. For simple queries only a list of column descriptions is shown directly beneath the query results. The descriptions are linked to the BioQ database documentation browser for more detailed information.
Detail View
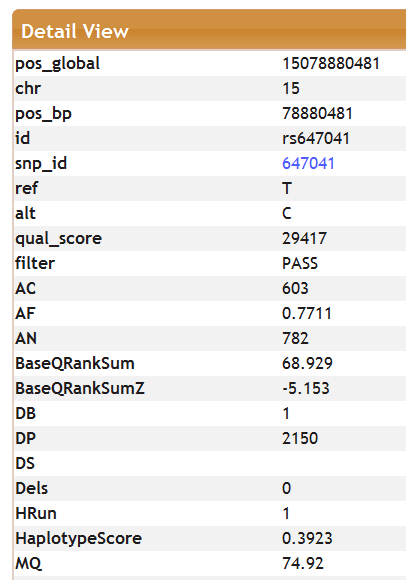
Selecting a row in the query results will open a new window below showing only that row in a separate table.
Linkage Disequilibrium (LD) Proxies
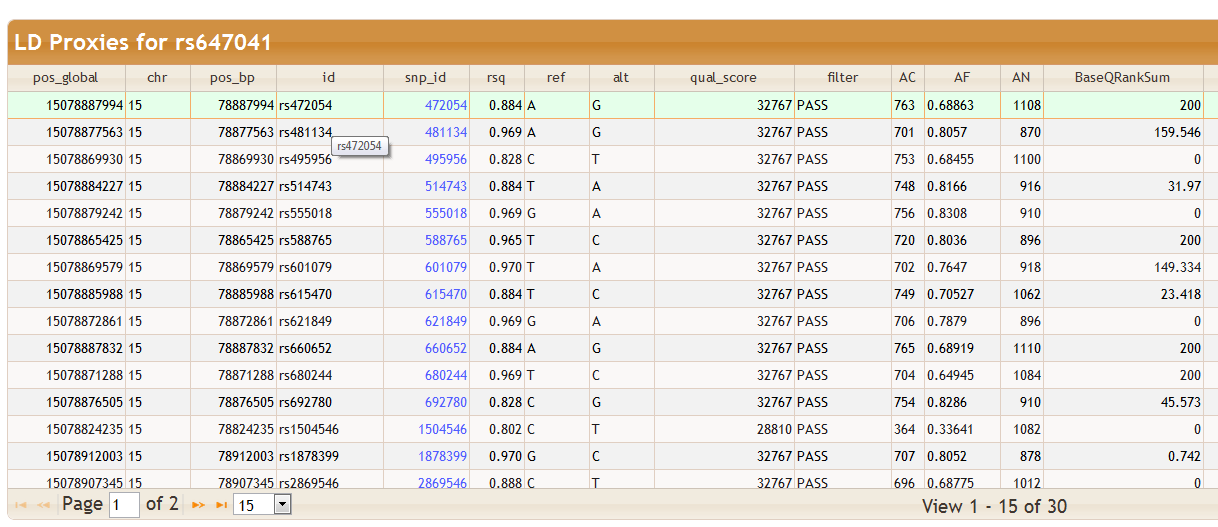
If the Add LD Proxies box directly above the query results is checked then, provided the query results contains a dbSNP ID column (usually named snp_id), a separate table showing all LD proxies will open up whenever a row is selected in the query results table. Here an LD proxy is defined as another SNP with r2 ≤ T where r is the correlation between SNPs measured in the selected population and T is the selected threshold. Note that the exact database and table used to measure LD is shown in the Program Parameters tab of the Additional Information section below: see the parameters bioq-ld-db and bioq-ld-table. In this case the table is bioq_hapmap_pr28_dbsnp132.ceu_plink_ld - this link takes you to the BioQ database documentation browser for information on how LD is estimated.
Downloads and Other Information
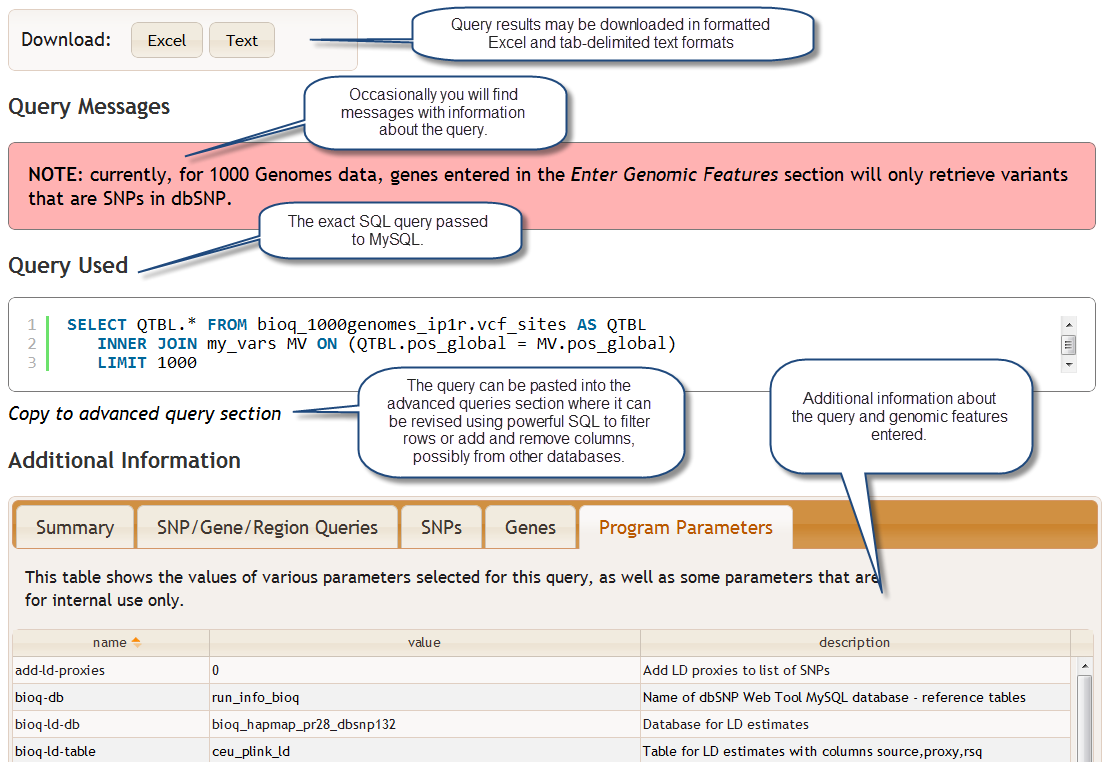
Beneath the query results you will find button for downloading the query results in formatted Excel and tab-delimited text formats. There is also a Query Messages section with detailed messages pertaining to your query, and a Query Used section that shows the SQL query that was passed to MySQL in order to generate the query results table. You can customize this query, and do things like filter the rows or add and remove columns, by clicking the text Copy to Advanced Query Section just below the Query Used section. See the section Advanced Queries with SQL for more information on using direct SQL queries. Finally, the Additional Information section provided a variety of details concerning the query and some diagnostic information about the software and what happened when the query was processed and run. You can find more information about this section in the section Querying Specific Genomic Features.