...
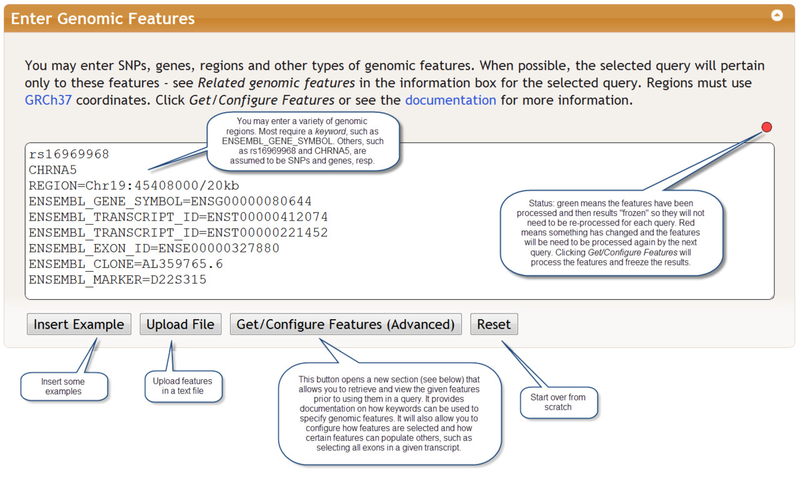
Features must be separated by carriage returns. They may be typed directly into the Enter Genomic Features section or uploaded as a file. An optional comment may be provided by inserting a comma after the feature followed by the comment. The status indicator indicator in the upper right of the features window turns green after a query and indicates that your features are now saved on the server. If you do not change these features, it will remain green and those features can be used for future queries. This saves the time of processing the features. The indicator turns red when you modify the queries. There is currently no undo mechanism, although your web browser may provide such a feature.
Subtopics
Entering Variants
dbSNP IDs
Specific genetic variants are entered dbSNP IDs. The "rs" (reference SNP) prefix is optional.
| Code Block |
|---|
|
[SNP=][rs]<numeric dbSNP ID> |
| Code Block |
|---|
| language | none |
|---|
| title | Examples |
|---|
|
16969968
rs16969968
SNP=16969968
SNP=rs16969968 |
Entering Genes
When a gene is specified, any SNP-related query will be applied to all SNPs found in this gene using the SNP/gene transcript relationships from the latest build of dbSNP (currently 134). Genes are specified using alphanumeric NCBI Entrez Gene symbols, like CHRNA5, or numeric Entrez Gene IDs, like 1138.
NCBI Entrez Gene Symbols (alphanumeric)
keyword: GENE
aliases: NCBI_GENE_SYMBOL, ENTREZ_GENE_SYMBOL, ENTREZ_GENE
A single gene may be specified using the NCBI Entrez Gene symbol. The GENE= prefix may optionally be used for clarity.
| Code Block |
|---|
|
[GENE=]<alphanumeric NCBI Entrez Gene symbol> |
| Code Block |
|---|
| language | none |
|---|
| title | Examples |
|---|
|
CHRNA5
GENE=CHRNA5 |
NCBI Entrez Gene IDs (numeric)
keyword: GENE_ID
aliases: NCBI_GENE_ID, ENTREZ_GENE_ID
Gene IDs must use the GENE_ID= prefix in order to distinguish them from dbSNP SNP IDs, which are the default numeric ID.
| Code Block |
|---|
|
GENE_ID=<numeric NCBI Entrez Gene ID> |
| Code Block |
|---|
| language | none |
|---|
| title | Examples |
|---|
|
GENE_ID=1138 |
Using search terms to specify genes
An entire set of genes may be specified using a the ENTREZ_GENE_QUERY= prefix. Multiple search terms may be grouped with the AND operator and phrases may be placed in double quotes.
| Code Block |
|---|
|
ENTREZ_GENE_QUERY="cholinergic receptor"
ENTREZ_GENE_QUERY=nicotinic AND beta AND muscle |
Entering Genomic Regions
keyword: REGION
Genomic regions are specified with a chromosome and starting and ending positions in base pairs.
| Code Block |
|---|
|
REGION=Chr<chromosome:1-22,X,Y>:<start position (bp)>..<end position (bp) |
| Code Block |
|---|
|
REGION=Chr15:78880000..78882000 |
Ensembl Features
| Info |
|---|
The Ensembl database is still in progress. |
Ensembl Clones
keyword: ENSEMBL_CLONE
A clone from the Ensembl Human Core database.
| Code Block |
|---|
|
ENSEMBL_CLONE=<alphanumeric clone ID> |
| Code Block |
|---|
|
ENSEMBL_CLONE=AL359765.6 |
The clone ID is matched against the column seq_region.name in the Ensembl Human Core database.
Ensembl Genes
keyword: ENSEMBL_GENE_SYMBOL
aliases: ENSEMBL_STABLE_GENE_ID
| Code Block |
|---|
|
ENSEMBL_GENE_SYMBOL=<alphanumeric stable Ensembl gene ID> |
| Code Block |
|---|
|
ENSEMBL_GENE_SYMBOL=ENSG00000080644 |
Ensembl Transcripts
keyword: ENSEMBL_TRANSCRIPT_ID
aliases: ENSEMBL_STABLE_TRANSCRIPT_ID
| Code Block |
|---|
|
ENSEMBL_TRANSCRIPT_ID=<alphanumeric stable Ensembl gene ID> |
| Code Block |
|---|
|
ENSEMBL_TRANSCRIPT_ID=ENST00000412074 |
...
Checking How BioQ Interpreted Your Features
After executing a query, at the bottom of the query results page you will find an Additional Information section. It provides some details on how BioQ processed the items you entered in the Genomic Features section. Suppose we entered the following features (note that we included optional comments followed by comma after each feature):
 Image Removed
Image Removed
The Summary tab provides some summary information about your features and will tell you if any errors were found.
 Image Removed
Image Removed
The SNP/Gene/Region Queries section displays the actual features, the optional comments, and how many variants and genes were retrieved for each features. It also shows the exact SQL WHERE statement that was used.
 Image Removed
Image Removed
The SNPs section displays the actual SNPs discovered from genomic features entered.
 Image Removed
Image Removed
The Genes section displays the genes resulting from the features entered.
...
Working with Features
When you click the Get/Configure Features button the following section appears:




