...
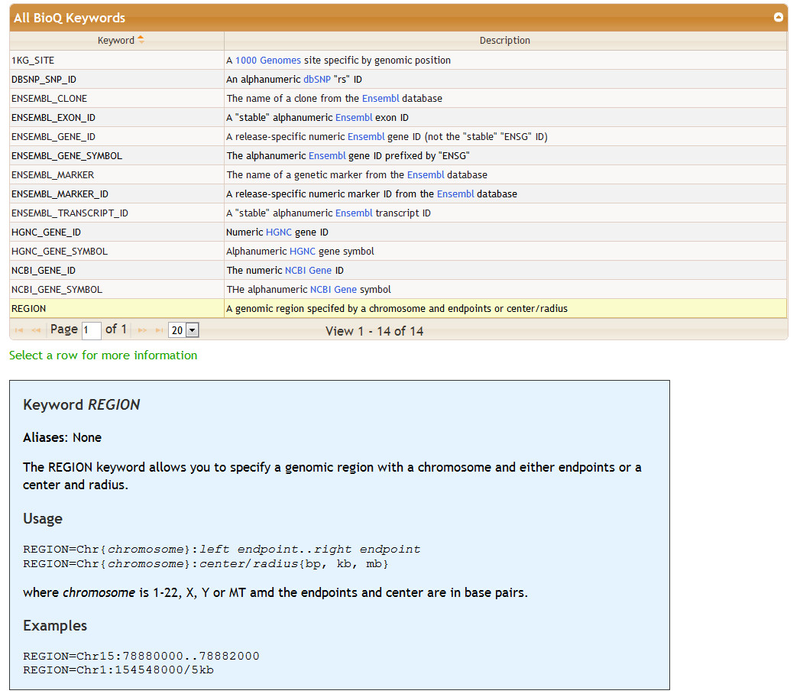
The bottom of the Genomic Features sectrion contains detailed information on how to specify features in the Enter Genomic Features section. Features are entered mainly by using keywords. A table of keywords is given - detailed information may be found by clicking on row. For some features the keywords are optional: for example, see the keywords DBSNP_SNP_ID and NCBI_GENE_SYMBOL.
Using Feature Tables in Queries
Some queries, such as the Genes query in the database Ensembl 64 - Human - Core, can be limited by certain feature tables:
If you do not enter any features, the query will retrieve a random set of data taken in whatever order the data exists in the database. If you enter features related to Ensembl Genes, such as by explicitly using the ENSEMBL_GENE_SYMBOL keyword or by using an interpopulation relationship such as getting the genes from regions, then the results of the Genes query will be limited to those features.
| Info |
|---|
The following is technical information on how feature tables are used in MySQL queries |
After the query is executed, if the query did use the Ensembl Genes feature table you will see a reference to its MySQL table feat_ensembl_gene in the Query Used section:
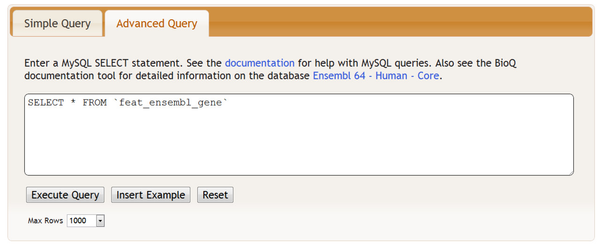
You may also access the feature tables in the Advanced Query section: