The Documentation Database
The BioQ web application uses several relational database models to provide documentation and query tools for the various genomic databases. All schemas and ER diagrams can be found in the MySQL Workbench document on our Subversion server.
Contents
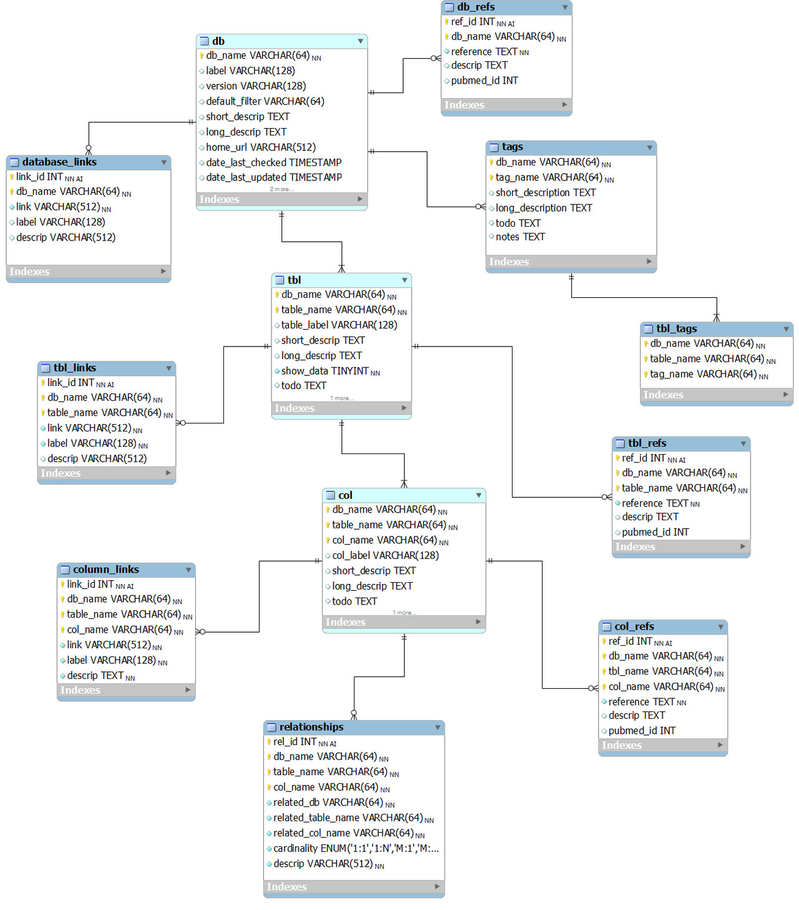
The core relational model
This entity-relationship (ER) diagram shows the core relational model for our database documentation (from subversion revision 70).
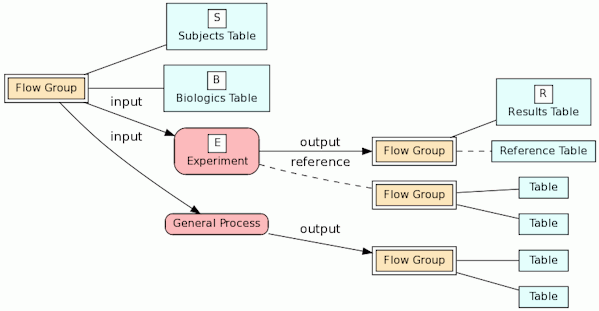
The biologic-experiment-result (BERT) relational model
Our BERT relational model is designed to document a genomic database by tracing the experimental source of the data. The key entities in the models are the biologics, the experiments and the experimental results as illustrated in the following diagram.
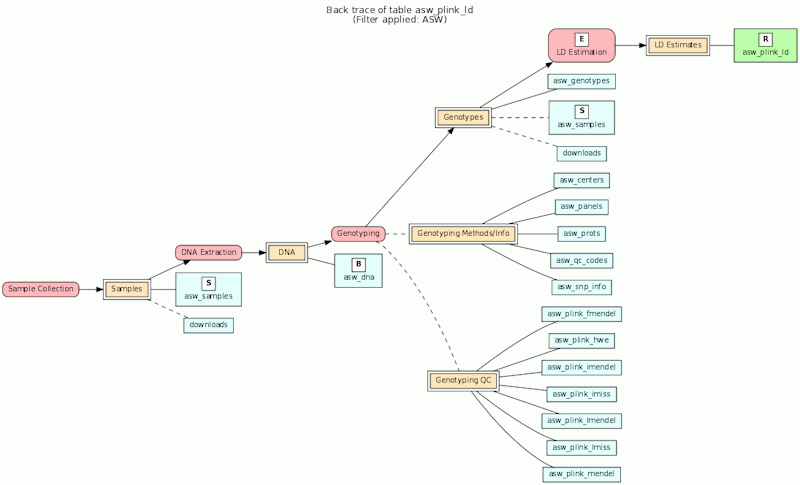
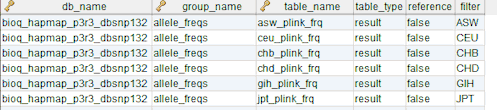
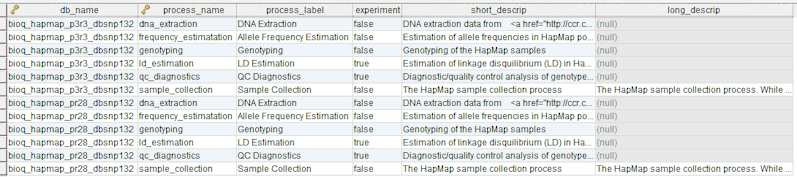
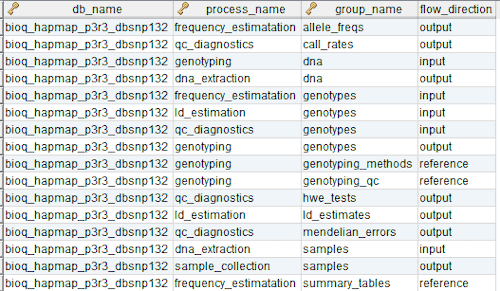
In the model, the tables in a relational database are grouped into flow groups that can input, output or reference material for the various experiments and processes that determined the data. Here is an example from our BioQ web application of the BERT model being applied to linkage disequilibrium (LD) data from the HapMap database. In BioQ, the diagram is interactive and can be used to navigate documentation and query the actual data.
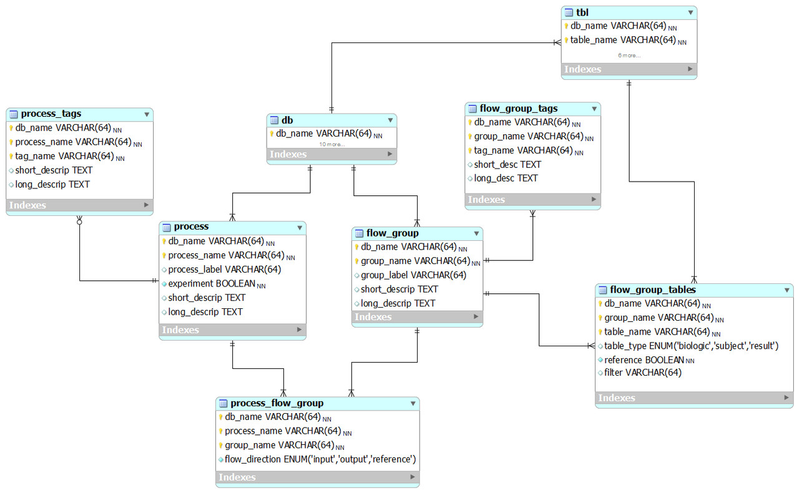
The following ER diagram shows the relational database implementation of the BERT model (from subversion revision 70, see the MySQL Workbench file ).
The queries model
This is our model for storing queries in the documentation database. This information is used to populate the BioQ queries for each database.
The features model
This is our model for querying genomic features such as genomic regions, gene transcripts, SNPs and other genetic variants. The way features are specified may depend on the database. For example, genes have different IDs in the NCBI, HUGO and Ensembl databases.
Features are specified in BioQ using keywords, such as SNP_ID and GENE_ID. There are usually defaults, such as a numeric feature being assumed to be a dbSNP rs ID and a alphanumeric string assumed to be a HUGO gene symbol.
Tables used for basic database documentation
db, tbl and col
These provide descriptions of database, tables and columns, respectively, in the BioQ genomic databases.
db_refs, tbl_refs and col_refs
Bibliographic references.
database_links, column_links, tbl_links
Web links.
tags
Tags are used to group tables into categories. The 'tags' table contains names and descriptions of the tags.
tbl_tags
This records which tags go with which tables.
Tables related to the BERT model and experimental process flow
flow_group
In the BERT model, a flow group is a group of tables that can be input or output for a specific process.
flow_group_tables
The tables in a flow group.
flow_group_tags
Tags that describe a flow group. These are not the same as filters.
process
A process in the BERT model.
process_flow_group
A group of tables, columns and/or databases which are input/output for a process or experiment in the BERT model. It's possible to have multiple groups with the same name for different processes. Each process must have a group and groups may be singletons.
process_tags
Tags that describe a process. These can also be used as filters. No entries as of version 70.
results_tables
For each table with table_type='result', determine if it can be traced to subjects and/or biologics.
Tables related to BioQ queries
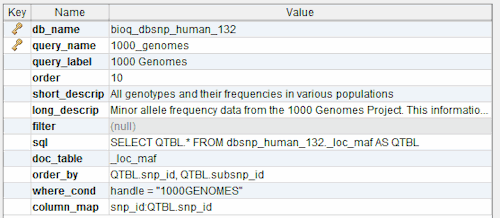
query
A query used in the BioQ web application.
query_column
This is used in Process.pm to look up information about columns being queried.
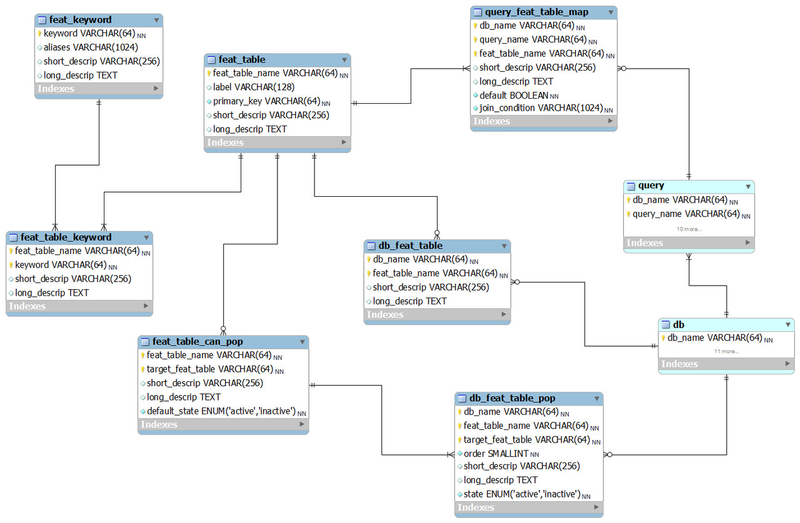
Tables related to genomic features
When querying databases in BioQ you may limit the queries to certain genomic features such as regions, genes and specific genetic variants. The way features are specified may depend on the database. For example, the same gene may have different IDs in the NCBI, HUGO and Ensembl databases. BioQ allows you to specifiy genomic features in a variety of ways. In some situations it will also add features automatically based on other features, such as finding all SNPs in a gene, and genes in a region, or all SNPs in a region. We are currently expanding this sytem to provider greater power in configuring how features are specified and how they relate to one another.
feat_keyword
A list of all the keywords and their aliases.
feat_table
The list of MySQL "feature tables" that contain feature data.
Some example of feature tables:
- feat_dbsnp_snp: a list of SNPs in the dbSNP database. Some keywords used to populate this table are DBSNP_ID (single SNP) and REGION (all SNPs in a given genomic region)
feat_table_keyword
This describes how specific feature tables are built using keywords. The keyword/feature table association is very important. It represents a process used to populate a feature table from a feature query (a feature specified on the BioQ query page). The descriptions in this table describe this process.
feat_table_can_pop
This explains how feature tables relate and interact with other feature tables. It contgains the other tables a feature table can populate. For example, the feature table feat_region can populate the table feat_dbsnp_snp via the relationship of the coordinate of a SNP being with the boundaries of a genomic region.
query_feat_table_map
This explains how to use feature tables once they are created. Specifically, it identifies the columns of a query that can be used to limit the query results by a feature table. For example, in the dbSNP database, the "SNP Summary" query can be limited to the features in the feat_dbsnp_snp feature table, as can any query that results in a dbSNP SNP ID column.
db_feat_table
The feature tables relevant to a specific database with descriptions of why the features are relevant and how they should be used.
db_feat_table_pop
This explains how specific databases use the feature tables and how they are inter-populated. For example, the 1000 Genomes databases probably should not populate gene transcript feature tables from genomic regions, although this behaviour may be configurable in future versions of BioQ. However, in 1000 Genomes we probably want the variant sites table feat_1kg_site to be populated from region features in feat_region.
Miscellaneous tables
reference
Reference table showing things like Subversion revision number.
relationships
For recording and describing relationships between tables in genomic databases. No entries as of version 70.
search_terms
Items used in the the dbDoc autocomplete search tool.
Updating the documentation schema
- Dump the existing database
- Use mysql --complete-insert --no-create-db --no-create-info --verbose
- See /projects/bioinf/ssaccone/dbdoc_util/dump_dbdoc_data.sh
- If necessary, create the schema file (current using MySQL Workbench with the file db_doc_model.mwb)
- The schema file is currently named db_doc_model.sql
- Note that the schema file should not have a 'USE' statement - instead use the dbdoc_util.pl dbdoc-db option.
- Run dbdoc_util.pl initdb
- You may also use the predefined script template_init.sh and the correpsonding options file template_init.opt
- Load the original dumped database
- See also the script /projects/bioinf/ssaccone/dbdoc_util/update_schema.sh
Initializing documentation for a single database
This is used to take an existing genomic database, such as HapMap, and set up entries in the dbDoc database for all the tables and columns in the database. This will ensure that entries for the different tables and columns exist so that more detailed documentation can be added.
Example: see /projects/bioinf/ssaccone/hapmap/dbdoc/hapmap_p3r3_dbsnp132.sh
- Run dbdoc_util.pl initdoc (with options) to set up dbDoc for a specified existing genomic MySQL database
- Run dbdoc_util.pl updatedoc --dbdoc-xml-file=<file> to read XML documentation
- Suggestion: break up XML documentation into multiple files, such as a "core" file that does not change often, and a another file that might describe what the recent changes are. This is currently done for the HapMap databases.
Modifying documentation
- The documentation for individual databases in the runk/main/databases directory of the Subversion repository contain shell scripts for running dbdoc_util.pl. These scripts read the XML documentation files to the build the MySQL documentaiton database
- Documentation for an entire databases may be deleted by deleting row for that databases from the table db via MySQL cascade.
Locations of Documentation on Subversion
- Documentation for individual databases is in the trunk/main/databases directory of the Subversion repository.
- Global documentation on BioQ genomic features, such as the list of feature keywords and tables, can be found in web/perl/dbdoc/features.xml